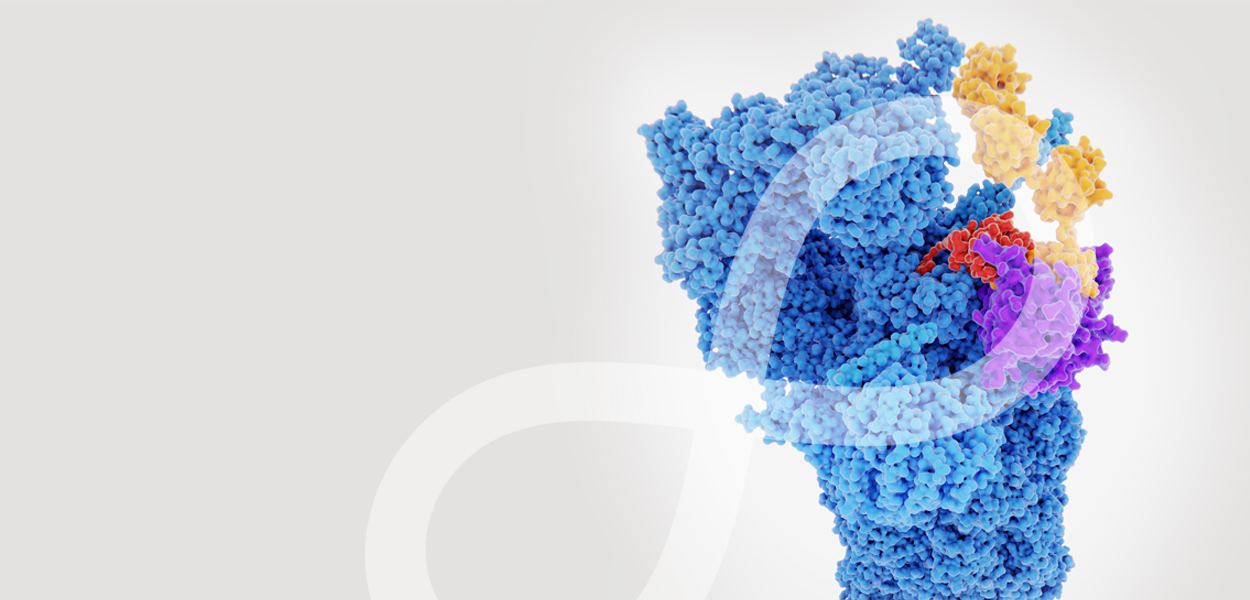
Panasas blends the superior performance and scalability of a parallel file system with the rock-solid resiliency of a cloud object store. The PanFS® parallel file system is the industry’s only solution with adaptable file-level erasure coding and comprehensive resiliency built into the core software. That means as you expand your operations, you don’t need to worry about data loss or service interruptions – reliability actually increases as you scale.
The only proven self-healing, self-managing parallel file system designed to accelerate HPC and AI applications with zero data loss, zero downtime, and zero headaches.
An entire storage lifecycle
No downtime
Exascale data environments
1 admin
Go from dock to data in
24 Hrs
Never compromise between performance and reliability again.
Trusted by global enterprise giants and research pioneers
Scale smarter, not harder.

Get the scalability, speed, and insights that your data demands without the associated complexity. Panasas solutions are so easy to deploy, operate, and maintain that a single IT admin with no specialized expertise can oversee a PanFS environment of any size. Experience the high-performance storage solution that has unparalleled reliability and simple manageability hardwired into its design DNA.
Get Started Today

1. Contact Us
Fill our contact form and schedule a consultation and a demo with our solutions experts.

2. Get a Cost Estimate
Based on the requirements, we share a proposal with timeline estimates suited to your infrastructure needs.

3. Kickoff
Starting with an infrastructure assessment, our engineers recommend tailored solutions.
SIGN-UP FOR A DEMO
Thank You
Thank you! We'll be in touch.